ExoPlaSim Python API Documentation
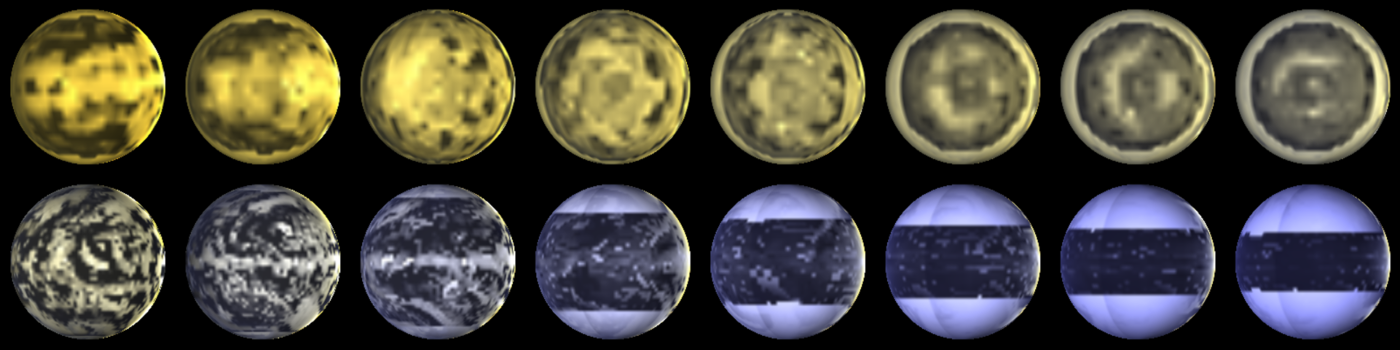
A range of planets modeled by ExoPlaSim, and postprocessed with SBDART. The top row consists of tidally-locked aquaplanets at T21 orbiting stars ranging from 2500 K to 4000 K, with orbital periods increasing with stellar mass. The bottom row consists of aquaplanets with 24-hour rotation at T42, orbiting stars ranging from 4000 K to 8000 K.
Contents
Created by Adiv Paradise
Copyright 2020, Distributed under the General Public License.
Requirements
Python 3.8+ (including development libraries, e.g. python-dev or python3.9-dev on Ubuntu–if using anaconda, these should already be included in your installation)
numpy
scipy (only needed for additional utilities, postprocessor)
GNU C (gcc/g++) and Fortran (gfortran) compilers (development headers must be present)
(optionally) Other compilers whose use you prefer for the model itself
(optionally) MPI libraries for those compilers
Compatibility
Linux (tested on Ubuntu 22.04): Yes
Google Colaboratory: Yes (note that OpenMPI support on Colaboratory is limited due to automatic root privileges; look up how to run OpenMPI executables with root permissions and note that this is not recommended)
Windows 10: Yes, via Windows Subsystem for Linux
Mac OS X: Yes, requires Xcode and developer tools, and OpenMPI support requires that Fortran-compatible libraries be built. Tested on Mac OS X Catalina and Big Sur (with MacPorts, GCC10, OpenMPI, and Anaconda3), Apple M1 compatibility has not been tested.
Optional Requirements
netCDF4 (for netCDF support)
h5py (for HDF5 support)
matplotlib (for plotting and interactivity with
makestellarspecandrandomcontinentsmodules)
Future Development Roadmap:
New features to be included in ExoPlaSim in the future include (but are not limited to) the following, in the order they are likely to appear:
Coupling with N-body integrators (confirmed for 4.0.0)
Support for an arbitrary number of light-sources in the sky, with individual temperatures and time-varying brightnesses, as well as single and compound eclipses/transits (confirmed for 4.0.0)
Support for time-varying geothermal/tidal heat flux into the atmosphere/ocean from the lithosphere (likely 4.0.0 or 4.1.0)
Coupling to the GENIE modeling framework to include a dynamic ocean and biogeochemistry
Support for extended light sources such as giant planets and rings
CO2 tracer transport and condensation
Support for swapping in a different radiation model
New in 3.4:
This is the final 3.x maintenance release before 4.0.0 (coming soon!).
Updated documentation
Several bugfixes including to core model operations such as the physics filters
Revamped
pyfftcompilation to eliminate thorny version dependencies–installation and compilation should be a lot smoother now.Modernized PlaSim source to be compliant with Fortran 2018 and MPI 4 (required for OpenMPI 5 and GCC 13)
Added pass-throughs to the MPI executable to allow runtime flags related to e.g. hyperthreading
New in 3.3:
New aerosol module for transport and gravitational settling of dust and photochemical hazes
New in 3.2:
Experimental integration with petitRADTRANS to compute transit spectra and reflectance spectra, including maps and true-colour images (use at your own risk)
Ability to specify general keplerian orbits, with high eccentricity, using a revamped orbit code for higher accuracy
Orbital elements now included in standard output
Numerous bugfixes
New in 3.0:
ExoPlaSim no longer depends on X11 libraries for installation and compilation!
Revamped postprocessor no longer depends on NetCDF-C libraries, and supports additional output formats (including netCDF, HDF5, NumPy archives, and archives of CSV files).
GCC and gfortran support through GCC 10.
Improved cross-platform compatibility
Numerous bugfixes
Installation
pip install exoplasim
The first time you import the module and try to create a model
after either installing or updating, ExoPlaSim will run a
configuration script and compile the pyfft library.
NEW in 3.2.2, updated in 3.4.0: If you need to re-run the configuration, because
for example the system libraries/compilers have changed, or the
configuration failed the first time (usually because OpenMPI
was not properly configured/visible), then you can call sysconfigure()
to rerun the configuration script.
You may also configure and compile the model manually if you wish
to not use the Python API, by entering the exoplasim/ directory
and running first configure.sh, then compile.sh (compilation flags
are shown by running ./compile.sh -h).
If there is a problem with gcc or gfortran being available, pyfft compilation may also fail
or need to be updated. To re-run the pyfft compilation, you can run compile_pyfft().
Both of these routines will be called the first time a Model is compiled, if they have not
ever been run before. You do not need to run these in order to start compiling and using the
model. They will also call each other if they detect that compiler detection has not yet been
completed and pyfft has not yet been compiled.
A clean install with conda:
ExoPlaSim requires at minimum a C compiler, a C++ compiler, and a Fortran compiler.
For parallel execution, an MPI compiler is also required.
It is heavily recommended that you use the GNU compilers and OpenMPI.
On linux that means gcc, gfortran, g++, and openmpi. On OS X it may be possible to use
other C and C++ compilers. Intel Fortran will work with the ExoPlaSim source, but not
with pyfft compilation through numpy, which means the postprocessor will not work
properly with Intel Fortran. That may or may not be a dealbreaker for you.
One of the easiest and safest ways to ensure you have met the compilation dependencies is to use something like conda. While ExoPlaSim itself does not (yet) have a compilation recipe available through conda or conda-forge, there are versions of gcc, gxx, gfortran, and openmpi that can be installed through conda via the conda-forge channel.
An ExoPlaSim installation (including compiler configuration and pyfft compilation) in a
conda environment would therefore look like:
conda install -c conda-forge gcc gxx gfortran "openmpi<5.0.1" python
pip install exoplasim
python -c "import exoplasim; exoplasim.sysconfigure()"
Note that at present, OpenMPI 5.0.1 onwards from conda-forge is affected by a critical bug related to linking against outdated C libraries on the conda-forge side; ExoPlaSim will fail to compile with OpenMPI if using conda-forge OpenMPI later than 5.0.0.
Most Common Error Modes
There are 3 major ways in which ExoPlaSim can crash. One is related to installation, one is related to model compilation/configuration, and one is related to numerical stability.
If in the run folder, diagnostic files are produced that appear to have made it all the way to the end of the year (there is a summary tag giving time elapsed and that sort of thing), then the problem is likely with the postprocessor. It is likely that the error output will be informative; if it is not clear how to resolve, please let me (the developer) know.
If the postprocessor itself is not the problem, then it’s likely you somehow passed incorrect output codes to the postprocessor. This is the most common scenario for postprocessor-related crashes. Check your inputs for any errors. In particular, note that climatology outputs are not available if storm climatology was not enabled.
If things crashed and burned immediately, it’s likely a configuration problem. There are two kinds of configuration problems that can commonly cause problems: a problem with how system libraries are configured, or a problem with how your model is configured.
If it appears that a file is missing or a command can’t be found:
This usually means a system library such as the Fortran compiler,
OpenMPI compiler, or numpy’s f2py utility is either not installed,
incorrectly installed, or not visible to ExoPlaSim. To verify, you can
run exoplasim.printsysconfig(ncpus=1) and exoplasim.printsysconfig(ncpus=2).
These commands (new in 3.2.2) print the contents of the single-core
and parallel executation configuration files, and additionally return their
contents as a python dictionary. There should not be any empty settings.
If something appears missing such as an MPI or Fortran compiler, and you
believe it’s installed, you can check by running e.g. mpifort --help
or gfortran --help. Additionally, for pyfft problems, verify that numpy’s
f2py utility is available by running python -c "import numpy.f2py". If any of these
fails but the library in question is installed, that suggests it is not in
the system path (the list of directories where programs may look for libraries
and executables). Ensure all libraries are properly installed, configured,
and on the path, then run sysconfigure() to reconfigure ExoPlaSim.
If it appears that the model has actually crashed:
Check to make sure you aren’t using a restart file from a run
that used a different resolution, or stellar spectrum files that aren’t
formatted correctly (use the makestellarspec
utility to format Phoenix spectra for ExoPlaSim), or boundary condition
.sra files that aren’t properly-formatted.
If things were fine until they weren’t, then it’s likely ExoPlaSim encountered a numerical instability of some kind. Some of these are physical (e.g. you ran a model at a thousand times Earth’s insolation, and the oceans boiled, or the model was too cold and the physics broke), while some are not (something happened to violate the CFL condition for the given timestep, or an unphysical oscillation wasn’t damped properly by the dynamical core and it grew exponentially). If this happens, either try a model configuration that is more physically reasonable, or if the problem appears not to have been physical, try reducing the timestep or increasing hyperdiffusion. Sometimes it also works to slightly adjust a model parameter such as surface pressure by a fraction of a percent or less–just enough to nudge the model out of whatever chaotic local minimum it ran into, but not enough to qualitatively change the resulting climate.
New in ExoPlaSim 3.0.0, there is a “crash-tolerant” run mode. With this mode enabled, a runtime crash will result in rewinding 10 years and resuming. This deals with many of the most frustrating problems related to numerical instability. However, due to the potential for infinite loops, this is only recommended for advanced users.
PlaSim Documentation
Original PlaSim documentation is available in the exoplasim/docs/ folder and online.
Usage
To use the ExoPlaSim Python API, you must import the module, create a Model or one of its subclasses, call its configure method and/or modify method, and then run it.
An IPython notebook is included with ExoPlaSim; which demonstrates basic usage. It can be found in the ExoPlaSim installation directory, or viewed directly here.
Basic example::
import exoplasim as exo
mymodel = exo.Model(workdir="mymodel_testrun",modelname="mymodel",resolution="T21",layers=10,ncpus=8)
mymodel.configure()
mymodel.exportcfg()
mymodel.run(years=100,crashifbroken=True)
mymodel.finalize("mymodel_output")
In this example, we initialize a model that will run in the directory
“mymodel_testrun”, and has the name “mymodel”, which will be used to
label output and error logs. The model has T21 resolution, or 32x64,
10 layers, and will run on 8 CPUs. By default, the compiler will use
8-byte precision. 4-byte may run slightly faster, but possibly at the
cost of reduced stability. If there are machine-specific optimization
flags you would like to use when compiling, you may specify them as a
string to the optimization argument, e.g. optimization='mavx'. ExoPlaSim
will check to see if an appropriate executable has already been created,
and if not (or if flags indicating special compiler behavior such as
debug=True or an optimization flag are set) it will compile one. We then
configure the model with all the default parameter choices, which means
we will get a model of Earth. We then export the model configurations
to a .cfg file (named automatically after the model), which will allow
the model configuration to be recreated exactly by other users. We
run the model for 100 years, with error-handling enabled. Finally, we
tell the model to clean up after itself. It will take the most recent
output files and rename them after the model name we chose, and delete
all the intermediate output and configuration files.
A Note on NetCDF and the (deprecated) Burn7 Postprocessor
As of ExoPlaSim 3.0.0, burn7 is deprecated. It is only
available via the exoplasim-legacy package.